Join GitHub today
GitHub is home to over 36 million developers working together to host and review code, manage projects, and build software together.
Sign upHow to retrieve all candidates in PubChem using MetFragR or MetFragCL #24
Comments
 JustinZZW
changed the title
JustinZZW
changed the title
How to retrieve all candidates using MetFragR or MetFragCL
How to retrieve all candidates in PubChem using MetFragR or MetFragCL
May 20, 2019
This comment has been minimized.
This comment has been minimized.
schymane
commented
May 20, 2019
|
I have example functions, data and documentation here: We are looking at improving this our side and streamlining documentation etc, if you can install this package from github this should get you started in the interim. The IPB team also have other workflows that may be useful for you. |
This comment has been minimized.
This comment has been minimized.
|
Thanks, Emma, It founds 1022 candidates, and return a final result with 682 candidates. However, which function should I use to export the all 1022 candidates? Or how do I modify the parameter to download all 1022 candidates? This is the log:
|
This comment has been minimized.
This comment has been minimized.
schymane
commented
May 20, 2019
|
The log indicates that your post processing settings are reducing the candidates: I highly recommend you choose some examples and try them on the web interface, see what settings result in the parameter files (which you can download) and use this to choose the options that suit what you want to do. |
This comment has been minimized.
This comment has been minimized.
|
Thanks, I can set "filter_by_InChIKey" and "filter_isotopes" as FALSE to turn off the post-filtering. However, it still indicates 70 cancidates were discarded before processing due to pre-filter. How do I turn off this function? Besides, I try the same netural mass in the web server with same setting, but it only retrieve 700 candidates. So I wonder to know why the difference? In the local computer, I use the MetFrag2.4.3-CL. I attach the config, log and config in webserver. |
This comment has been minimized.
This comment has been minimized.
schymane
commented
May 20, 2019
|
These candidates are salts / mixtures / disconnected and cannot possibly be observed at the mass of interest and are thus excluded entirely. It is a factor of the way the data is retrieved from PubChem. Since they will not be observed at the mass you have given, it makes no sense to include them, so we have not added an "on/off" option for this case. |
This comment has been minimized.
This comment has been minimized.
schymane
commented
May 20, 2019
|
To check compatibility between web and CL version in detail, you have to compare the parameter files. Some discrepancy can arise if the web uses the local PubChem mirror versus the live online PubChem query. |
This comment has been minimized.
This comment has been minimized.
|
Thanks a lot for the patient explanation. I agree your point that the difference may be casued by the pubchem mirror and online query. But how can I know which it used in webserver and CL verison? I compared both parameter files, and do not find the corresponding parameter to clarify. In addition, if the CL verison use the online query, how to make the result reproducible? Thanks again. |
This comment has been minimized.
This comment has been minimized.
schymane
commented
May 20, 2019
|
You have to check the database parameters to see. On the web, if you tick the references box, it automatically uses the online version. I can't seem to download your parameter file from here. I just gave and posted a talk on this today, maybe the slides help a little? https://zenodo.org/record/3046373 |
This comment has been minimized.
This comment has been minimized.
|
Thanks very much for the slides. It's really useful and impressive. First, if the webserver use the online verison, I confused why it retrive less compounds (700 cmps) than the CL verison (1022 cmps)? Second, I'm not very clear what's the meaning of "filter_isotopes"? In the help file, it said it remove all candidates containing non-standard isotopes. May you help to give some examples? I really appreicate your kindness reply. This is parameters of the CL version:
This is parameters of the webserver:
|
This comment has been minimized.
This comment has been minimized.
schymane
commented
May 21, 2019
|
I am not sure what the problem was with the file download, it redirected me to a completely different website instead of the parameter file, but it may have been the app I was using. Many thanks for copying the parameters, this is very helpful. Also just to clarify: the CL uses the online version (unless you specify a local version of PubChem), the web uses an offline version by default, but when I select the references, I see now MetFragDatabaseType = ExtendedMetChem Please let me know if this resolves the issue. If you still see different candidate numbers after testing this, can you submit your MetFrag web parameters to the team in Halle so they can investigate further? |
This comment has been minimized.
This comment has been minimized.
|
Thanks, I tried as your instructions with 4 different parameter sets. It still have large differnces in candidatdes in CL (1022) and webserver (700). The filter_isotopes leads only 1 candidate difference. Therefore, as you say, I guess the webserver use the offline version, but the CL use the online version. Following are results from 4 groups:
|
This comment has been minimized.
This comment has been minimized.
schymane
commented
May 21, 2019
|
@sneumann @korseby can you confirm that MetFrag web only works on an offline mirror now (is there any way to set it to use the online version instead)? Does this explain the candidate difference? What is the date of the PubChem mirror? This is a huge difference in candidate numbers ... quite surprising. Thanks! |
This comment has been minimized.
This comment has been minimized.
|
Hi, I can confirm that the online version is using an older local mirror of PubChem. |
This comment has been minimized.
This comment has been minimized.
This comment has been minimized.
This comment has been minimized.
|
I noticed that you use an outdated version of the MetFrag-CLI. It is also possible to use the local MetChem mirror (deployed as docker container) from the command line (which speeds up processing). Please also make sure to have the latest version installed (currently 2.4.5). The easiest way would be to install it from bioconda, see here: https://github.com/bioconda/bioconda-recipes/tree/bc656c004ca3399595959403ca1870e83ac1e50b/recipes/metfrag or to use the biocontainers image. There is also a command line reference for MetFrag-CLI, see here: http://ipb-halle.github.io/MetFrag/projects/metfragcl/ |
This comment has been minimized.
This comment has been minimized.
|
ok, I will try the lastest version. Thanks! @korseby |
JustinZZW commentedMay 20, 2019
Hi,
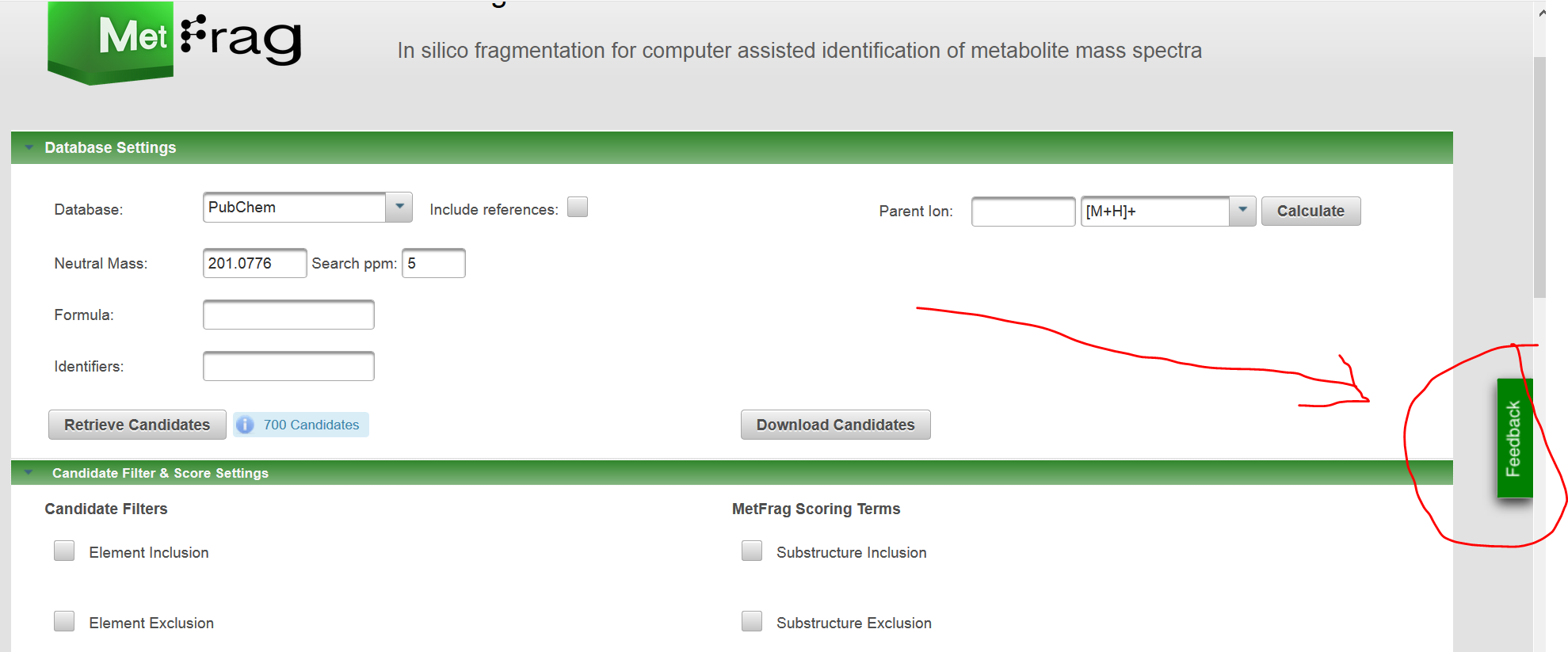
I wonder to know how to retrieve all possible candidates in PubChem with defined NeutralPrecursorMass and DatabaseSearchRelativeMassDeviation using MetFragR? I noticed in CASMI 2017, it said "The candidates were retrieved as InChI structures from PubChem (mirror dated 2017-02-03) using MetFrag 2.4.2". It is easily completed in the MetFrag webserver, but I do not how to do it in MetFragR or MetFragCL.
Thanks very much!